Detectmulticollinearity¶
Tabular Data - Data EDA - Correlation - Detect Multicollinearity¶
Table of Content¶
How to detect correlation between features?¶
Correlation is measured between numeric features only.
Starting with calculating the correlation metric between numeric features
corr_matrix = df.select_dtypes('number').corr()
Secondly, visualizing the metric using heatmap
_, ax = plt.subplots(figsize =(10,8)) sns.heatmap(corr_matrix,ax = ax, linewidths = 0.1) plt.title('Features Correlation') plt.savefig('../report/plots/correlation.jpg') plt.show()
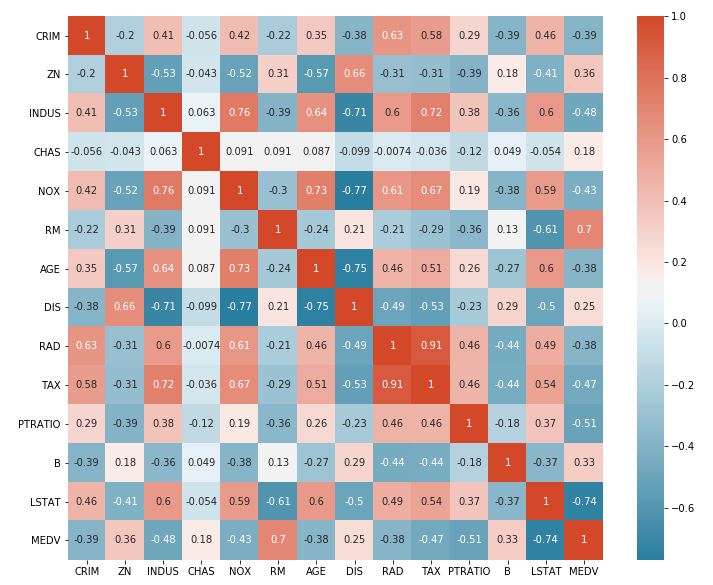
Heatmap is not very easily to be visualized, So for better interpreting, we will jump to third step.
Thirdly, Calculating the correlation between each feature and all the other features.
# Simplify by emptying all the data below the diagonal # np.tril_indices_from will return the indecies diagonal and below (lower triangle) tril_index = np.tril_indices_from(corr_matrix) # Make the unused values NaNs for coord in zip(*tril_index): corr_matrix.iloc[coord[0], coord[1]] = np.NaN # Stack the data and convert to a dataframe corr_values = (corr_matrix .stack() .to_frame() .reset_index() .rename(columns={'level_0':'feature1', 'level_1':'feature2', 0:'correlation'})) # Get the absolute values for sorting corr_values['abs_correlation'] = corr_values.correlation.abs()
This will output a dataframe, containing each feature and its correlation coefficient with other features.
feature1
feature2
correlation
abs_correlation
f1
f2
-0.52
0.52
f1
f3
0.89
0.89
f2
f3
-0.023
0.023
Fourthly, Ploting a histplot to get an overview of the features correlation.
ax = corr_values.abs_correlation.hist(bins=50, figsize=(12, 8)) ax.set(xlabel='Absolute Correlation', ylabel='Frequency')
Lastly, Check which of the featueres has a correlation above certain threshold like 0.8
corr_values.sort_values('correlation', ascending=False).query('abs_correlation>0.8')